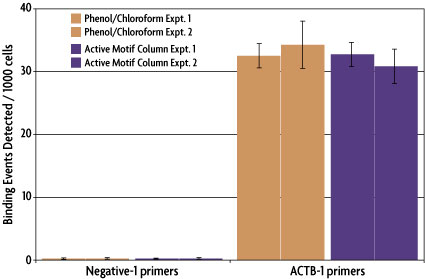
Magna ChIRP RNA Interactome Kit -Isolation and characterization of non-coding RNA:chromatin complexes | 17-10494

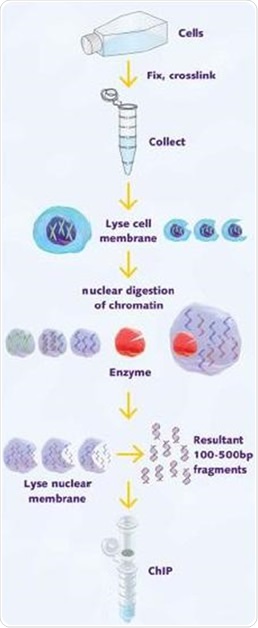
Purification procedure. (A) Diagram of chromatin purification and in... | Download Scientific Diagram
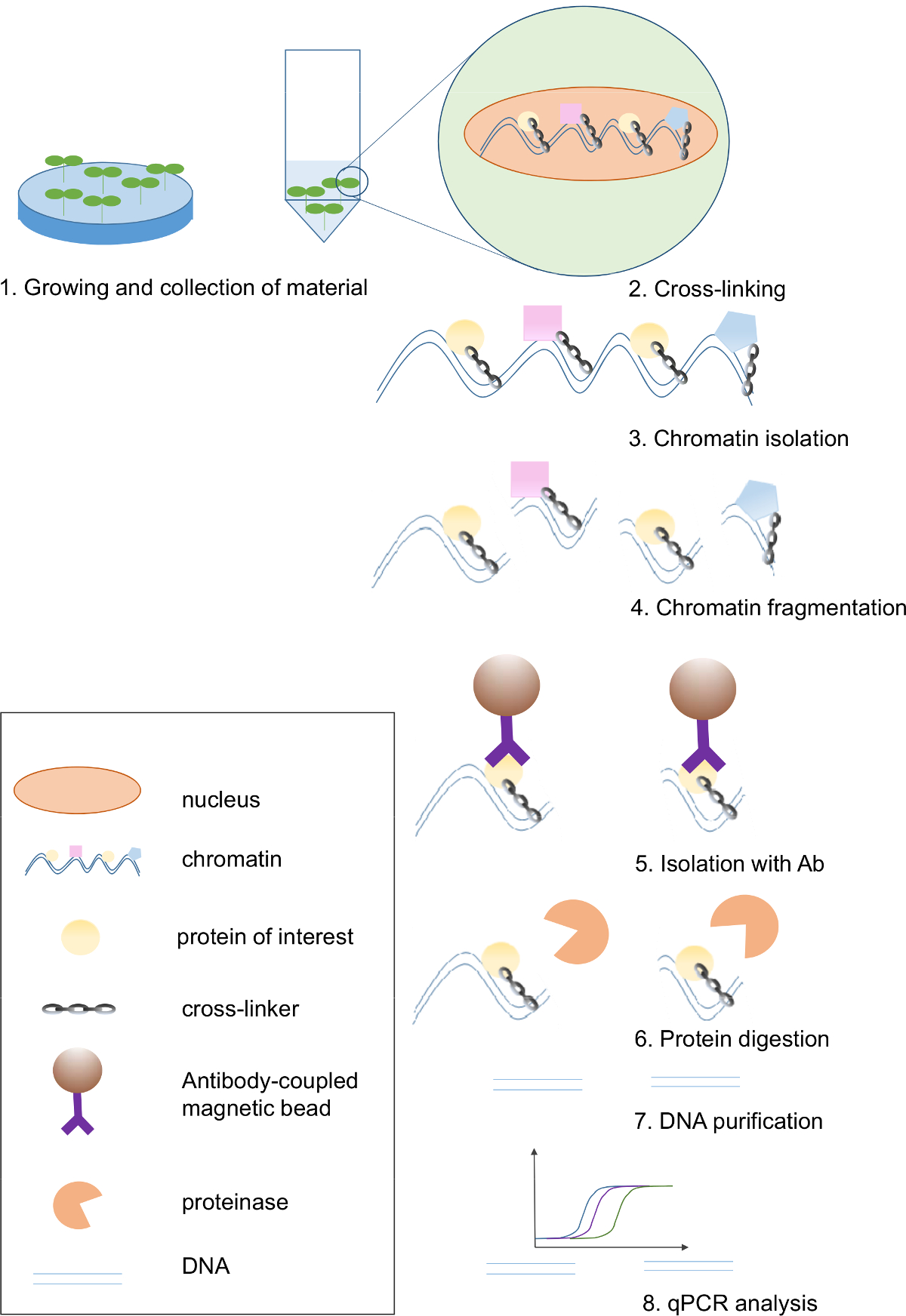
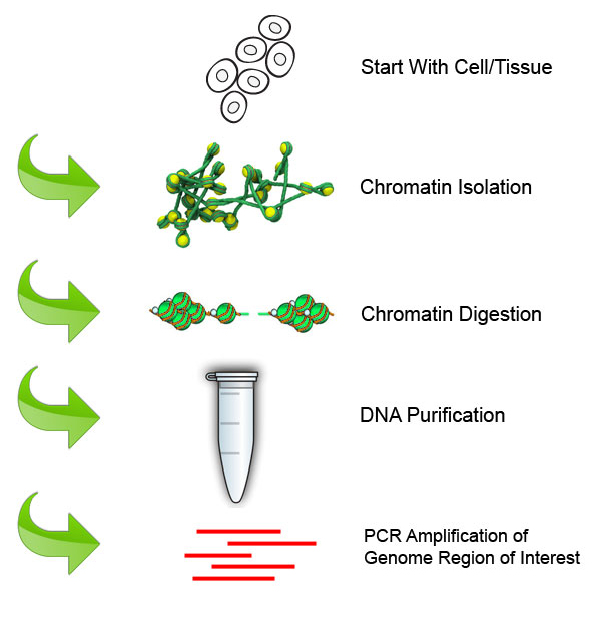
Chromatin Immunoprecipitation Assay for the Identification of Arabidopsis Protein-DNA Interactions In Vivo
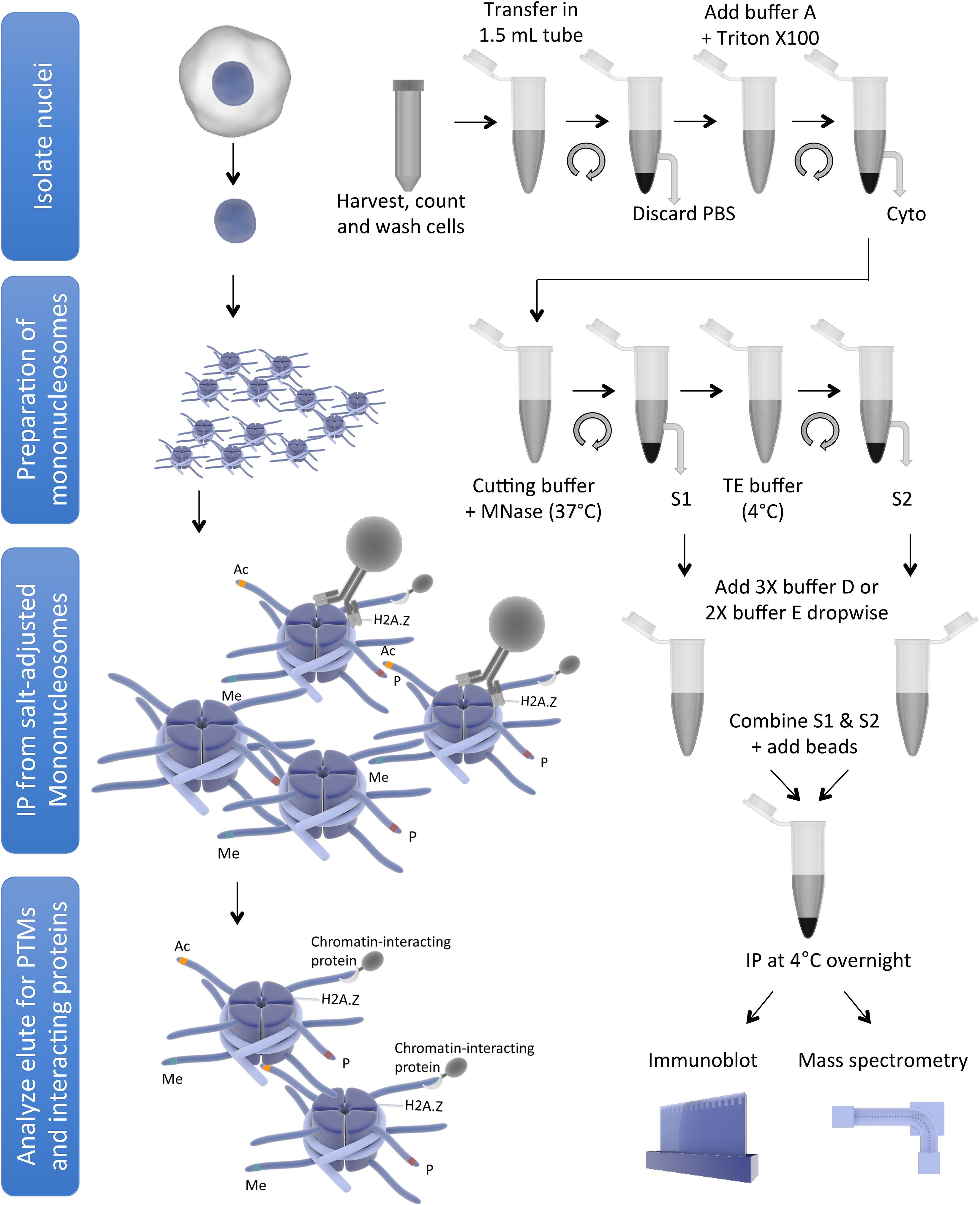
Frontiers | The Use of Mononucleosome Immunoprecipitation for Analysis of Combinatorial Histone Post-translational Modifications and Purification of Nucleosome-Interacting Proteins
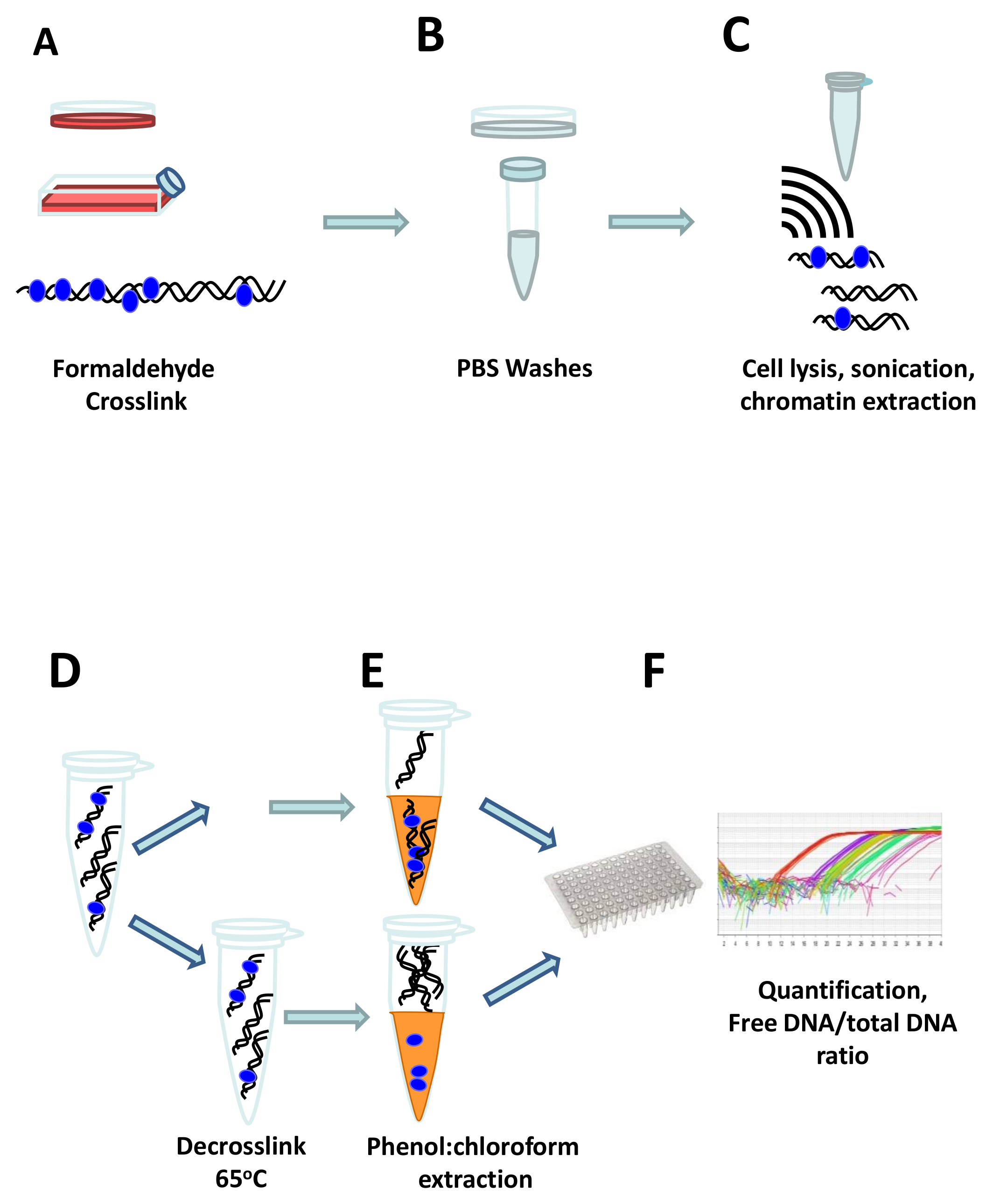
Formaldehyde-assisted Isolation of Regulatory Elements to Measure Chromatin Accessibility in Mammalian Cells | Protocol (Translated to French)
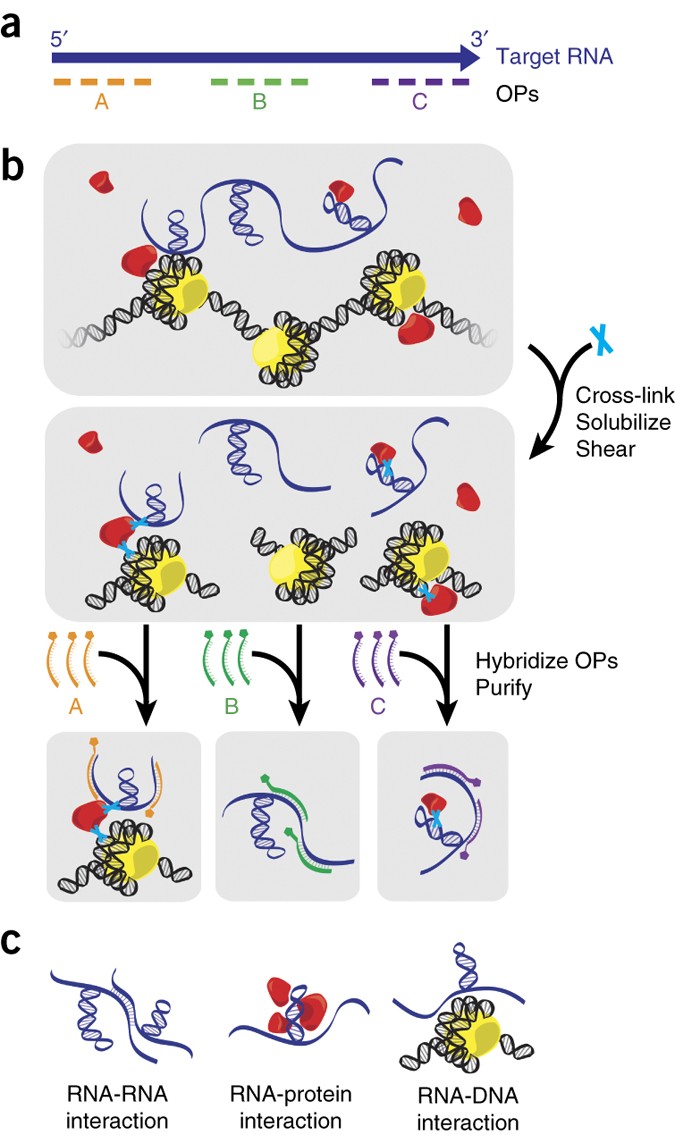
Revealing long noncoding RNA architecture and functions using domain-specific chromatin isolation by RNA purification | Nature Biotechnology
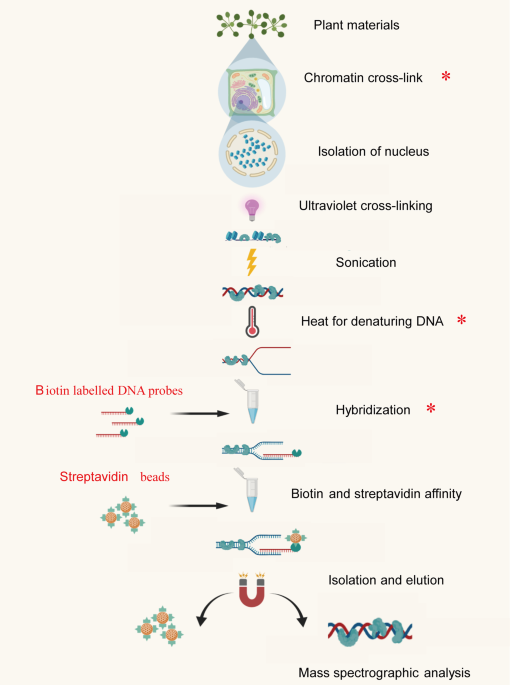
Reverse Chromatin Immunoprecipitation (R-ChIP) enables investigation of the upstream regulators of plant genes | Communications Biology

Efficient sequence‐specific isolation of DNA fragments and chromatin by in vitro enChIP technology using recombinant CRISPR ribonucleoproteins - Fujita - 2016 - Genes to Cells - Wiley Online Library

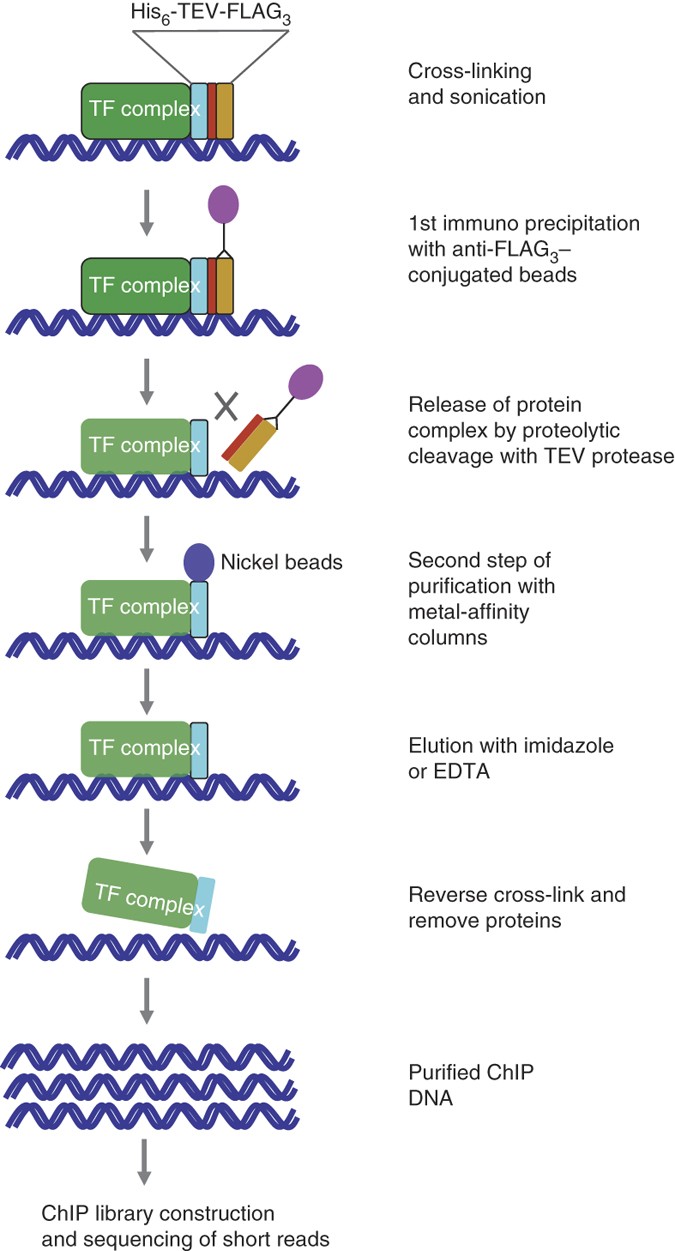
Protocol for fractionation-assisted native ChIP (fanChIP) to capture protein-protein/DNA interactions on chromatin - ScienceDirect
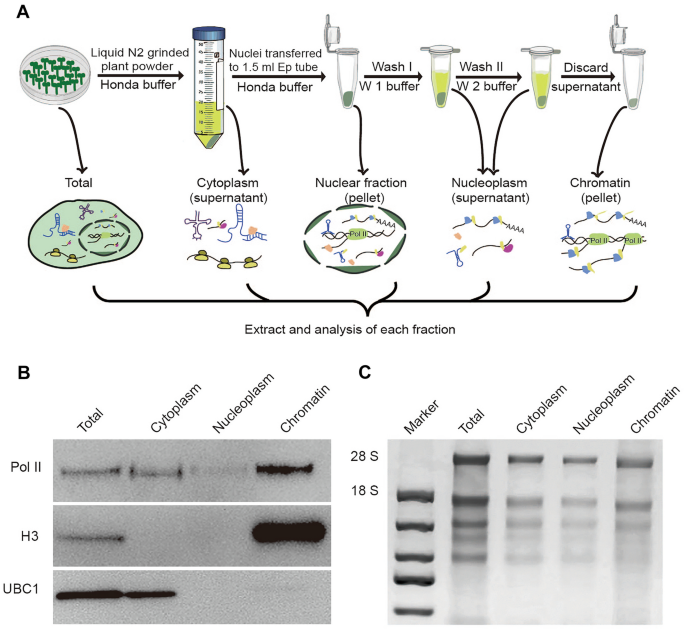
A simple and robust method for isolating and analyzing chromatin-bound RNAs in Arabidopsis | Plant Methods | Full Text
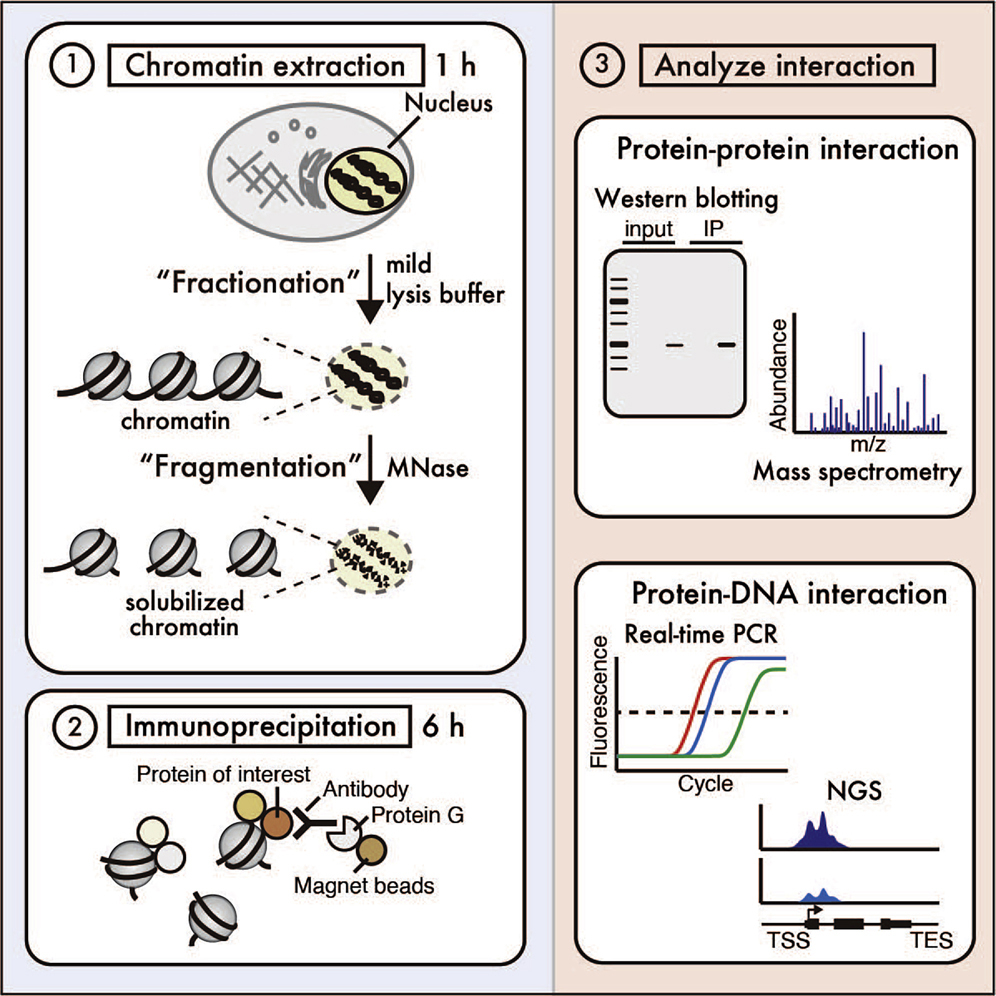




![PDF] Chromatin Isolation by RNA Purification (ChIRP) | Semantic Scholar PDF] Chromatin Isolation by RNA Purification (ChIRP) | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/971ae2b323439a21f4be734a4ebd953d5cb3f7cd/4-Figure1-1.png)